apply flashr with OCV
wei wang
Last updated: 2018-02-11
Code version: dcd6356
preparation
- creat a folder
mkdir Breastcancer- put your data into this folder
here we use the file ‘example.mat’. the reason we use .mat file is because there is a matlab package in our comparision.
- creat the wrapper function in R and matlab
wrapper function for OCV
CVPMD_softImpute=function(Y,c_s,K,fold = 10, method = "PMD"){
N = dim(Y)[1]
P = dim(Y)[2]
colindex = matrix(sample(P,P),ncol = fold)
rowindex = matrix(sample(N,N),ncol = fold)
missing= array(0,dim = c(fold,N,P))
foldindex = array(0,dim = c(fold,fold,2))
for(i in 1:fold){
for(j in 1:fold){
foldindex[i,j,1] = i
foldindex[i,j,2] = (i+j) %% fold
}
}
foldindex[which(foldindex == 0)] = fold
for(i in 1:fold){
missing[i, , ] = Y
for(j in 1:fold){
missing[i,rowindex[,foldindex[j,i,1]],colindex[,foldindex[j,i,2]]] = NA
}
missing[i,,which(colSums(missing[i,,],na.rm = T) ==0)] = Y[,which(colSums(missing[i,,],na.rm = T) ==0)]
}
# c_s is the candicate of shrinkage parameter
n_s = length(c_s)
# rmse for each grids
CVRMSE = rep(0,n_s)
minrmse = Inf
opt_s = 0
# for each candidate, we run it N_sim times
for(t_s in 1:n_s){
# for each grid
# each time we set the rmse to zeros
rmse = rep(0,fold)
for(i in 1:fold){
if(method == "PMD"){
res_log = capture.output({out = PMD(missing[i,,], sumabs = c_s[t_s], sumabsv = NULL, sumabsu = NULL,K = K)})
}else{
out = softImpute(missing[i,,], rank.max = K,lambda = c_s[t_s])
}
if(length(out$d)==1){
misshat = (out$d) * out$u %*% t(out$v)
}else{
misshat = out$u %*% diag(out$d) %*% t(out$v)
}
for(j in 1:fold){
# for each fold j
rmse[i] = rmse[i] + sum((Y[rowindex[,foldindex[j,i,1]],colindex[,foldindex[j,i,2]]] -
misshat[rowindex[,foldindex[j,i,1]],colindex[,foldindex[j,i,2]]])^2,na.rm = TRUE)
}
} #get the result for one run
CVRMSE[t_s] = CVRMSE[t_s] + sqrt(sum(rmse)/(N*P))
if(CVRMSE[t_s] < minrmse){
minrmse = CVRMSE[t_s]
opt_s = c_s[t_s]
}
}
return(list(opt_s = opt_s, output = CVRMSE))
}
PMA.wrapper = function(Y,ngrids = 10,K,fold = 10){
library(PMA)
N = dim(Y)[1]
P = dim(Y)[2]
c_s = seq(0.3,0.8,len=ngrids)
cvout = CVPMD_softImpute(Y,c_s,K ,fold , method = "PMD")
res_log = capture.output({out = PMD(Y,sumabsu = NULL, sumabsv = NULL, sumabs = cvout$opt_s ,K = K)})
return(list(d = out$d, u = out$u, v = out$v))
}
softImpute.wrapper = function(Y,ngrids = 10,K,fold = 10){
library(softImpute)
N = dim(Y)[1]
P = dim(Y)[2]
c_s = seq(0,100,len=ngrids)
cvout = CVPMD_softImpute(Y,c_s,K ,fold , method = "softImpute")
out = softImpute(Y, rank.max = K,lambda = cvout$opt_s)
return(list(d = out$d, u = out$u, v = out$v))
}
OCV_index=function(Y,k_fold = 5){
N = dim(Y)[1]
P = dim(Y)[2]
colindex = matrix(sample(P,P),ncol = k_fold)
rowindex = matrix(sample(N,N),ncol = k_fold)
foldindex = array(0,dim = c(k_fold,k_fold,2))
for(i in 1:k_fold){
for(j in 1:k_fold){
foldindex[i,j,1] = i
foldindex[i,j,2] = (i+j) %% k_fold
}
}
foldindex[which(foldindex == 0)] = k_fold
return(list(foldindex = foldindex, rowindex = rowindex, colindex = colindex))
}
# OCVindex = OCV_index(Y,k_fold = 5)
OCV_data = function(Y,OCVindex,k_fold = 5){
N = dim(Y)[1]
P = dim(Y)[2]
colindex = OCVindex$colindex
rowindex = OCVindex$rowindex
foldindex = OCVindex$foldindex
missing= array(0,dim = c(k_fold,N,P))
for(i in 1:k_fold){
missing[i, , ] = Y
for(j in 1:k_fold){
missing[i,rowindex[,foldindex[j,i,1]],colindex[,foldindex[j,i,2]]] = NA
}
missing[i,,which(colSums(missing[i,,],na.rm = T) ==0)] = Y[,which(colSums(missing[i,,],na.rm = T) ==0)]
}
return(missing)
}
# OCVdata = OCV_data(Y,OCVindex,k_fold = 5)
OCV_SSE = function(Y,OCVindex,OCVdata,k_fold = 5,method = "flash",Kmax = 50){
colindex = OCVindex$colindex
rowindex = OCVindex$rowindex
foldindex = OCVindex$foldindex
missing = OCVdata
SSE = rep(0,k_fold)
for(i in 1:k_fold){
miss_hat = call_method(missing[i,,], method = method, Kmax = Kmax)
for(j in 1:k_fold){
SSE[i] = SSE[i] + sum((Y[rowindex[,foldindex[j,i,1]],colindex[,foldindex[j,i,2]]] -
miss_hat[rowindex[,foldindex[j,i,1]],colindex[,foldindex[j,i,2]]])^2,na.rm = TRUE)
}
}
RMSE = sqrt(sum(SSE)/(dim(Y)[1] * dim(Y)[2]))
return(RMSE)
}
call_method = function(Y_data,method,Kmax = 50){
if(method == "flash"){
data = flashr::flash_set_data(Y_data)
f_greedy = flashr::flash_add_greedy(data,K=Kmax)
Y_hat = f_greedy$EL %*% t(f_greedy$EF)
}else if(method == "flash_wn"){
data = flashr::flash_set_data(Y_data)
f_greedy = flashr::flash_add_greedy(data,K=Kmax,nullcheck=FALSE)
Y_hat = f_greedy$EL %*% t(f_greedy$EF)
}else if(method == "flash_gb"){
data = flashr::flash_set_data(Y_data)
f_greedy = flashr::flash_add_greedy(data,K=Kmax)
f = flashr::flash_backfit(data,f_greedy)
Y_hat = f$EL %*% t(f$EF)
}else if(method == "flash_bf"){
data = flashr::flash_set_data(Y_data)
f_data = flashr::flash_add_factors_from_data(data,K = Kmax)
f = flashr::flash_backfit(data,f_data)
Y_hat = f$EL %*% t(f$EF)
}else if(method == "PMD"){
res_log = capture.output({out = PMA.wrapper(Y_data,ngrids = 10,K = Kmax,fold = 10)})
if(length(out$d)==1){
Y_hat = (out$d) * out$u %*% t(out$v)
}else{
Y_hat = out$u %*% diag(out$d) %*% t(out$v)
}
}else if(method == 'PN'){
library(ebnm)
library(flashr)
data = flashr::flash_set_data(Y_data)
f_greedy = flashr::flash_add_greedy(data,K=Kmax,ebnm_fn = ebnm_pn)
Y_hat = f_greedy$EL %*% t(f_greedy$EF)
}else if(method == 'PMD_d'){
out = PMA::PMD(Y_data, K = Kmax)
Y_hat = out$u %*% diag(out$d) %*% t(out$v)
}else if(method == 'softImpute_d'){
gsoft = softImpute::softImpute(Y_data, rank.max = Kmax)
Y_hat = gsoft$u %*% diag(gsoft$d) %*% t(gsoft$v)
}else if(method == "softImpute"){
gsoft = try(softImpute.wrapper(Y_data,ngrids = 10,K = Kmax,fold = 10))
if(length(gsoft$d)==1){
Y_hat = try((gsoft$d) * gsoft$u %*% t(gsoft$v))
}else{
Y_hat = try(gsoft$u %*% diag(gsoft$d) %*% t(gsoft$v))
}
}else if(method == "SF_flash"){
gsoft = try(softImpute.wrapper(Y_data,ngrids = 10,K = Kmax,fold = 10))
LL = gsoft$u %*% diag(gsoft$d)
FF = gsoft$v
data = flashr::flash_set_data(Y_data)
f_data = flashr::flash_add_lf(data, LL=LL, FF=FF)
f = flashr::flash_backfit(data,f_data)
Y_hat = f$EL %*% t(f$EF)
}else{
# stop("the method is not implemented yet, please check it out")
Y_hat = matrix(0,ncol = dim(Y_data)[2],nrow = dim(Y_data)[1])
}
return(Y_hat)
}
NSF_OCV = function(OCVindex){
if(file.exists("NBSFAout")){
unlink("NBSFAout", recursive= T)
}
system("mkdir NBSFAout")
# write the missing index
writeMat("./NBSFAout/OCVindex.mat", OCVindex = OCVindex)
system("matlab -nosplash -nodesktop -r \"addpath(\'../\'); run(\'missvalue.m\');exit;\" ")
g_nsfa = readMat("./NBSFAout/NSFAresult.mat")
nsfa_rmse = as.vector(g_nsfa$RMSE)
return(nsfa_rmse)
}This file name is wrapper.R
matlab code
In this folder
ddpath('~/HG/flash/data/missingvalue/methods/NBSF/nsfa-master/');
addpath('~/HG/flash/data/missingvalue/methods/NBSF/nsfa-master/utils/');
% you need to change the path and the data file (not centered)
load ../Ydata.mat;
Ycentered=Y;
settings=defaultsettings();
[settings.D,settings.N]=size(Ycentered);
settings.iterations=100;
% now the missing is decide by the this is the missing index.(or you can upload the missing matrix)
file_path = fullfile(pwd,'NBSFAout/OCVindex.mat');
load(file_path);
% missindex = ones(settings.D,settings.N); this is not missing index this
% is observed index.
missindex = (~isnan(Y));
k_fold = 10;
foldindex = zeros(k_fold,k_fold,2);
for k = 1:2
for j = 1:k_fold
for i = 1:k_fold
foldindex(i,j,k) = OCVindex.foldindex(i + k_fold*(j - 1) + k_fold * k_fold * (k - 1));
end
end
end
SSE = zeros(1,k_fold);
for i = 1:k_fold
mvmask = missindex;
for j = 1:k_fold
mvmask(OCVindex.rowindex(:,foldindex(j,i,1)),OCVindex.colindex(:,foldindex(j,i,2))) = 0;
end
mvmask(:,sum(mvmask) == 0) = (missindex(:,sum(mvmask) == 0));
initialsample=init_nsfa(settings);
[finalsample,resultstable]=nsfa(Ycentered,mvmask,initialsample,settings);
Y_hat = (finalsample.G * finalsample.X);
for j = 1:k_fold
SSE(i) = SSE(i) + ...
sum(nansum((Ycentered(OCVindex.rowindex(:,foldindex(j,i,1)),OCVindex.colindex(:,foldindex(j,i,2)))- ...
Y_hat(OCVindex.rowindex(:,foldindex(j,i,1)),OCVindex.colindex(:,foldindex(j,i,2)))).^2));
end
end
RMSE = sqrt(sum(SSE)/(settings.D*settings.N));
RMSE;
save_path = fullfile(pwd,'NBSFAout/NSFAresult');
save(save_path,'RMSE');you need change the path ~/HG/flash/data/missingvalue/methods/NBSF/nsfa-master/ and ~/HG/flash/data/missingvalue/methods/NBSF/nsfa-master/utils/ which have the NBSFA matlab code.
creat the run.R file to run the code
In this case, we have problem in running flashr on RCC. so we just try other methods first.
source("../wrapper.R")
library(R.matlab)
## run the code
Y_centered = readMat("../example.mat")
Y = Y_centered$Y
# in the matlab package of NSF, the use the centered data by rows
N = dim(Y)[1]
P = dim(Y)[2]
Y = Y - rowMeans(Y) %*% t(rep(1,P))
OCVindex = OCV_index(Y,k_fold = 10)
OCVdata = OCV_data(Y,OCVindex,k_fold = 10)
PN_rmse = OCV_SSE(Y,OCVindex,OCVdata,k_fold = 10,method = "PN",Kmax = 40)
flashG_rmse = OCV_SSE(Y,OCVindex,OCVdata,k_fold = 10,method = "flash",Kmax = 40)
flashGwn_rmse = OCV_SSE(Y,OCVindex,OCVdata,k_fold = 10,method = "flash_wn",Kmax = 40)
flashB_rmse = OCV_SSE(Y,OCVindex,OCVdata,k_fold = 10,method = "flash_bf",Kmax = 40)
pmd_rmse = OCV_SSE(Y,OCVindex,OCVdata,k_fold = 10,method = "PMD",Kmax = 40)
soft_rmse = OCV_SSE(Y,OCVindex,OCVdata,k_fold = 10,method = "softImpute",Kmax = 40)
nsfa_rmse = NSF_OCV(OCVindex)
result = c(PN_rmse,flashG_rmse,flashGwn_rmse,flashB_rmse,nsfa_rmse,pmd_rmse,soft_rmse)
saveRDS(result, "./NBSFAout/output.rds")
print(sessionInfo())this file name is runOCV.R
To read different data sets, please click here for more details
creat folder to track the result
mkdir outlogcreat .sbatch file
#!/bin/bash
#SBATCH --job-name=arrayJob
#SBATCH --output=./outlog/arrayJob_%A_%a.out
#SBATCH --error=./outlog/arrayJob_%A_%a.err
#SBATCH --array=1-20
#SBATCH --time=26:00:00
#SBATCH --partition=mstephens
#SBATCH --ntasks=1
#SBATCH --mem-per-cpu=8000
######################
# Begin work section #
######################
# Print this sub-job's task ID
module load R
module load matlab
mkdir test${SLURM_ARRAY_TASK_ID}
cd test${SLURM_ARRAY_TASK_ID}
Rscript --verbose ../runOCV.Rthis file is name as OCV.sbatch
run OCV
now in your folder, you should have:
[weidong@midway2-login1 Breastcancer]$ ls
OCV.sbatch example.mat missvalue.m outlog runOCV.R wrapper.Rsbatch OCV.sbatchto get the result
T = 20
results = matrix(NA,ncol = 7, nrow = T)
for(i in 1:T){
test_folder = paste("test", i, sep = "")
out_folder = "NBSFAout"
out_file = "output.rds"
file_name = file.path(test_folder,out_folder,out_file)
results[i,] = try(readRDS(file_name))
}
saveRDS(results,"./boxplot.rds")plot the result
# this is the independent result
library(ggplot2)
plot_res = function(output,title = "data",legend_position = "none", x_label, myColors){
rmse = as.vector(output)
N = dim(output)[1]
methods = rep(x_label, each = N)
df = data.frame(RMSE = rmse, Method = methods )
p<-ggplot(df, aes(x=Method, y=RMSE, color=Method)) +
geom_boxplot()+
# geom_violin()+
ggtitle(title) + theme_bw()+ scale_color_manual(values=myColors)+
theme(legend.position= legend_position, legend.text=element_text(size=15),
plot.title = element_text(size = 15, face = "bold"),
axis.text.y = element_text(size =12.6),
axis.text.x = element_text(size =12.6,angle = 45, hjust = 1))
p
}
setwd("~/HG/flash_workflow/analysis")
fill_color = c("red","yellow3", "tan1","springgreen3", "springgreen","deepskyblue3","deepskyblue")
############## MovieLens data
ML_res = readRDS("../data/output/missingdata/MovieLens/ML100K_box.rds")
ML_res[c(2,13,17,21,29,37,62,76,77,93,95,100),] = NA
ML_res = matrix(as.numeric(ML_res),ncol = 4)
# c("flash","NBSFA","PMD","softImpute")
ml_res = readRDS("../data/output/missingdata/MovieLens/boxplot_cv.rds")
# flashG_rmse,flashGwn_rmse,pmd_rmse,soft_rmse
ml_res[1:20,] = sqrt((ml_res[1:20,])^2 * (943*1682)/(100000))
ml_res = ml_res[1:20,]
ML_res = ML_res[-c(2,13,17,21,29,37,62,76,77,93,95,100),]
ML_res = ML_res[1:20,]
# add PN result
ML_pn_res = readRDS("../data/output/missingdata/MovieLens/box_cv_pn.rds")
# the result order would be flashG PMD PMDcv NSFA SF SFcv
output = cbind(ML_pn_res[,1],ML_pn_res[,2],ML_res[,3],ml_res[,3],ML_res[,2],ML_res[,4],ml_res[,4])
x_label= c("flash","flash.pn","PMD","PMD.cv1","NBSFA","SI","SI.cv")
plotM = plot_res(output,"MovieLens data",x_label = x_label,myColors = fill_color)
############ Breast Cancer data
BC_res = readRDS("../data/output/missingdata/BreastCancer/BreastCancer_box.rds")
BC_res = BC_res[1:20,]
# c("flash","NBSFA","PMD","softImpute")
bc_res = readRDS("../data/output/missingdata/BreastCancer/box_Breast.rds")
# c("PN","flashG","flashGwn","flashB","nsfa","pmd","soft")
# the result order would be flashG PMD PMDcv NSFA SF SFcv
output = cbind(bc_res[,2],bc_res[,1],BC_res[,3],bc_res[,6],BC_res[,2],BC_res[,4],bc_res[,7])
x_label= c("flash","flash.pn","PMD","PMD.cv1","NBSFA","SI","SI.cv")
plotB = plot_res(output,"Breast cancer data",x_label = x_label,myColors = fill_color)
############ GTEx data
GZ_res = readRDS("../data/output/missingdata/GTExZsocre/gtexzscore_box.rds")
GZ_res = GZ_res[1:20,]
# c("flash","NBSFA","PMD","softImpute")
gz_res = readRDS("../data/output/missingdata/GTExZsocre/box_cv.rds")
# flashG_rmse,nsfa_rmse,pmd_rmse,soft_rmse
gz_pn_res = readRDS("../data/output/missingdata/GTExZsocre/box_pn.rds")
gz_pn_res[c(23,25,29,34),] = NA
gz_pn_res = matrix(as.numeric(gz_pn_res[21:40,]),ncol = 2)
# the result order would be flashG PMD PMDcv NSFA SF SFcv
output = cbind(gz_res[,1],gz_pn_res[,2],GZ_res[,3],gz_res[,3],GZ_res[,2],GZ_res[,4],gz_res[,4])
x_label= c("flash","flash.pn","PMD","PMD.cv1","NBSFA","SI","SI.cv")
plotG = plot_res(output,"GTEx data",x_label = x_label,myColors = fill_color)
############ Text data
PT_res = readRDS("../data/output/missingdata/DenoiseRtext/president_box.rds")
PT_res = PT_res[1:20,]
# c("flash","NBSFA","PMD","softImpute")
pt_res = readRDS("../data/output/missingdata/DenoiseRtext/box_president.rds")
# c("PN","flashG","flashGwn","flashB","nsfa","pmd","soft","zero")
# the result order would be flashG PMD PMDcv NSFA SF SFcv
output = cbind(pt_res[,3],pt_res[,1],PT_res[,3],pt_res[,6],PT_res[,2],PT_res[,4],pt_res[,7])
x_label= c("flash","flash.pn","PMD","PMD.cv1","NBSFA","SI","SI.cv")
plotP = plot_res(output,"Presidential address data",x_label = x_label,myColors = fill_color)
############ Tumor data
DT_res = readRDS("../data/output/missingdata/DenoiseRtumor/denoiseRtumor_box.rds")
DT_res = DT_res[1:20,]
# c("flash","NBSFA","PMD","softImpute")
dt_res = readRDS("../data/output/missingdata/DenoiseRtumor/box_denoiseTumor.rds")
# c("PN","flashG","flashGwn","flashB","nsfa","pmd","soft")
# the result order would be flashG PMD PMDcv NSFA SF SFcv
output = cbind(dt_res[,2],dt_res[,1],DT_res[,3],dt_res[,6],DT_res[,2],DT_res[,4],dt_res[,7])
x_label= c("flash","flash.pn","PMD","PMD.cv1","NBSFA","SI","SI.cv")
plotT = plot_res(output,"Tumor data",x_label = x_label,myColors = fill_color)# gridExtra::grid.arrange(plotP,plotT,plotG,plotB,plotM, layout_matrix = rbind(c(1,NA,2),c(NA,5,NA),c(4,NA,3)))
gridExtra::grid.arrange(plotM,plotT,plotG,plotB,plotP, layout_matrix = rbind(c(1,2,3),c(4,5,NA) ))Warning: Removed 4 rows containing non-finite values (stat_boxplot).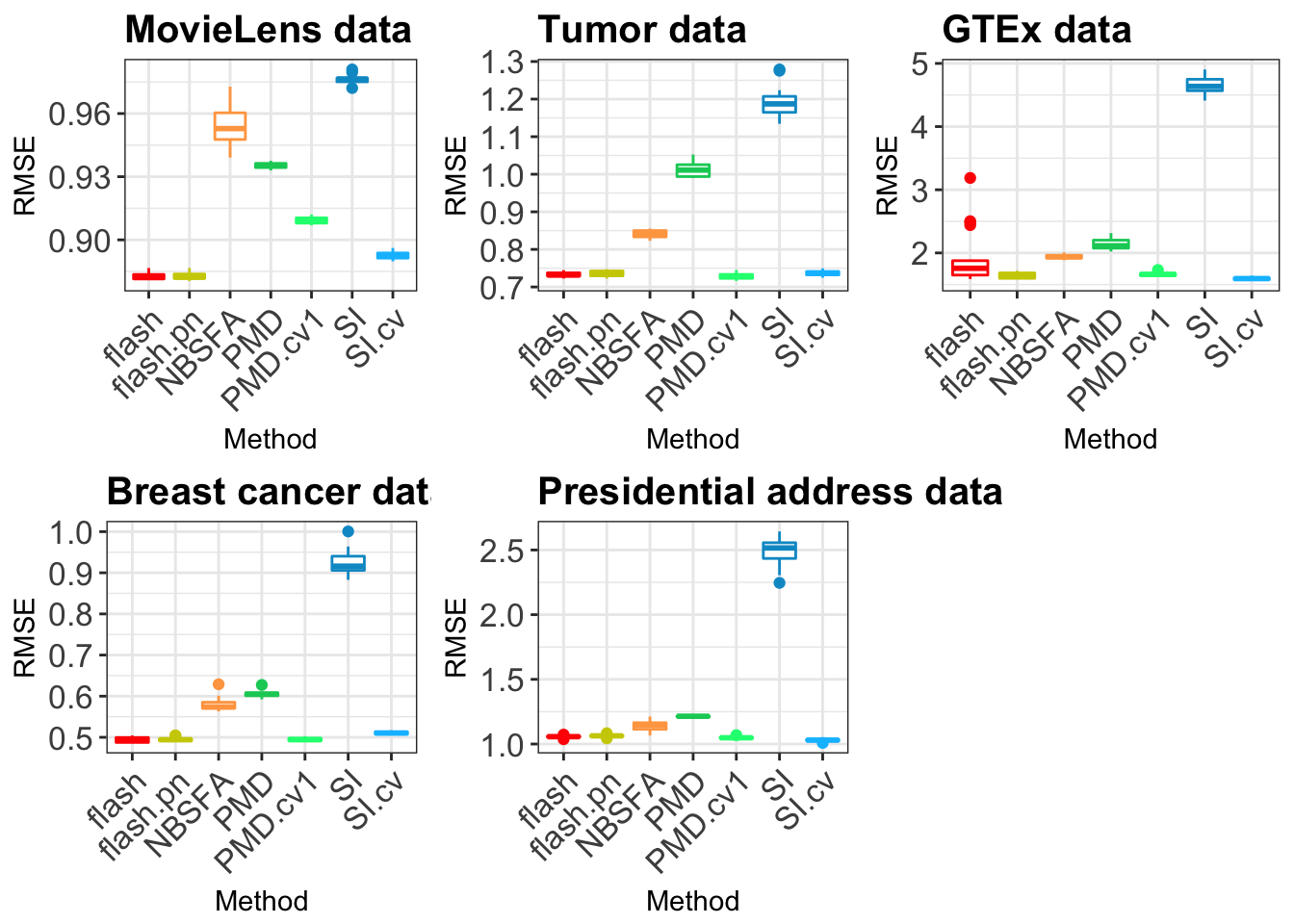
other information
please click here to check other information Missing data OCV settings
Session information
sessionInfo()R version 3.3.0 (2016-05-03)
Platform: x86_64-apple-darwin13.4.0 (64-bit)
Running under: OS X 10.13.3 (unknown)
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] denoiseR_1.0 scales_0.4.1 MASS_7.3-47 reshape2_1.4.3
[5] flashr_0.4-6 workflowr_0.4.0 rmarkdown_1.6 ggplot2_2.2.1
[9] R.matlab_3.6.1 softImpute_1.4 Matrix_1.2-11 PMA_1.0.9
[13] impute_1.48.0 plyr_1.8.4 ssvd_1.0
loaded via a namespace (and not attached):
[1] ashr_2.2-3 lattice_0.20-35 colorspace_1.3-2
[4] htmltools_0.3.6 yaml_2.1.16 rlang_0.1.6
[7] R.oo_1.21.0 withr_2.1.1 R.utils_2.5.0
[10] foreach_1.4.4 stringr_1.2.0 munsell_0.4.3
[13] gtable_0.2.0 R.methodsS3_1.7.1 devtools_1.13.3
[16] codetools_0.2-15 leaps_3.0 evaluate_0.10.1
[19] memoise_1.1.0 labeling_0.3 knitr_1.18
[22] pscl_1.5.2 doParallel_1.0.11 irlba_2.2.1
[25] parallel_3.3.0 curl_2.8.1 Rcpp_0.12.14
[28] flashClust_1.01-2 backports_1.1.2 scatterplot3d_0.3-40
[31] truncnorm_1.0-7 gridExtra_2.3 digest_0.6.13
[34] stringi_1.1.6 flashr2_0.4-0 grid_3.3.0
[37] rprojroot_1.2 tools_3.3.0 magrittr_1.5
[40] lazyeval_0.2.0 tibble_1.3.4 cluster_2.0.6
[43] FactoMineR_1.36 SQUAREM_2017.10-1 httr_1.3.0
[46] rstudioapi_0.6 iterators_1.0.9 R6_2.2.2
[49] git2r_0.19.0 This R Markdown site was created with workflowr